 |
LArSoft
v10_06_00
Liquid Argon Software toolkit - https://larsoft.org/
|
 |
LArSoft
v10_06_00
Liquid Argon Software toolkit - https://larsoft.org/
|
NeutrinoIdTool class. More...
#include "NeutrinoIdTool.h"
Classes | |
| class | SliceFeatures |
| Slice features class. More... | |
Public Member Functions | |
| NeutrinoIdTool () | |
| Default constructor. More... | |
| void | SelectOutputPfos (const pandora::Algorithm *const pAlgorithm, const SliceHypotheses &nuSliceHypotheses, const SliceHypotheses &crSliceHypotheses, pandora::PfoList &selectedPfos) |
| Select which reconstruction hypotheses to use; neutrino outcomes or cosmic-ray muon outcomes for each slice. More... | |
Private Types | |
| typedef std::pair< unsigned int, float > | UintFloatPair |
| typedef std::vector< SliceFeatures > | SliceFeaturesVector |
Private Member Functions | |
| void | GetSliceFeatures (const NeutrinoIdTool *const pTool, const SliceHypotheses &nuSliceHypotheses, const SliceHypotheses &crSliceHypotheses, SliceFeaturesVector &sliceFeaturesVector) const |
| Get the features of each slice. More... | |
| bool | GetBestMCSliceIndex (const pandora::Algorithm *const pAlgorithm, const SliceHypotheses &nuSliceHypotheses, const SliceHypotheses &crSliceHypotheses, unsigned int &bestSliceIndex) const |
| Get the slice with the most neutrino induced hits using Monte-Carlo information. More... | |
| bool | PassesQualityCuts (const pandora::Algorithm *const pAlgorithm, const float purity, const float completeness) const |
| Determine if the event passes the selection cuts for training and has the required NUANCE code. More... | |
| void | Collect2DHits (const pandora::PfoList &pfos, pandora::CaloHitList &reconstructedCaloHitList, const pandora::CaloHitSet &reconstructableCaloHitSet) const |
| Collect all 2D hits in a supplied list of Pfos and push them on to an existing hit list, check so not to double count. More... | |
| unsigned int | CountNeutrinoInducedHits (const pandora::CaloHitList &caloHitList) const |
| Count the number of neutrino induced hits in a given list using MC information. More... | |
| int | GetNuanceCode (const pandora::Algorithm *const pAlgorithm) const |
| Use the current MCParticle list to get the nuance code of the neutrino in the event. More... | |
| void | SelectAllPfos (const pandora::Algorithm *const pAlgorithm, const SliceHypotheses &hypotheses, pandora::PfoList &selectedPfos) const |
| Select all pfos under the same hypothesis. More... | |
| void | SelectPfosByProbability (const pandora::Algorithm *const pAlgorithm, const SliceHypotheses &nuSliceHypotheses, const SliceHypotheses &crSliceHypotheses, const SliceFeaturesVector &sliceFeaturesVector, pandora::PfoList &selectedPfos) const |
| Select pfos based on the probability that their slice contains a neutrino interaction. More... | |
| void | SelectPfos (const pandora::PfoList &pfos, pandora::PfoList &selectedPfos) const |
| Add the given pfos to the selected Pfo list. More... | |
| pandora::StatusCode | ReadSettings (const pandora::TiXmlHandle xmlHandle) |
Private Attributes | |
| bool | m_useTrainingMode |
| Should use training mode. If true, training examples will be written to the output file. More... | |
| std::string | m_trainingOutputFile |
| Output file name for training examples. More... | |
| bool | m_selectNuanceCode |
| Should select training events by nuance code. More... | |
| int | m_nuance |
| Nuance code to select for training. More... | |
| float | m_minPurity |
| Minimum purity of the best slice to use event for training. More... | |
| float | m_minCompleteness |
| Minimum completeness of the best slice to use event for training. More... | |
| float | m_minProbability |
| Minimum probability required to classify a slice as the neutrino. More... | |
| float | m_defaultProbability |
| Default probability set if score could not be calculated. More... | |
| unsigned int | m_maxNeutrinos |
| The maximum number of neutrinos to select in any one event. More... | |
| bool | m_persistFeatures |
| If true, the mva features will be persisted in the metadata. More... | |
| T | m_mva |
| The mva. More... | |
| std::string | m_filePathEnvironmentVariable |
| The environment variable providing a list of paths to mva files. More... | |
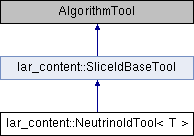
NeutrinoIdTool class.
Compares the neutrino and cosmic hypotheses of all of the slices in the event. Uses an MVA to calculate the probability of each slice containing a neutrino interaction. The N slices with the highest probabilities are identified as a neutrino (if sufficiently probable) all other slices are deemed cosmogenic.
If training mode is switched on, then the tool will write MVA training exmples to the specified output file. The events selected for training must pass (user configurable) slicing quality cuts. Users may also select events based on their interaction type (nuance code).
Definition at line 33 of file NeutrinoIdTool.h.
|
private |
Definition at line 163 of file NeutrinoIdTool.h.
|
private |
Definition at line 162 of file NeutrinoIdTool.h.
| lar_content::NeutrinoIdTool< T >::NeutrinoIdTool | ( | ) |
Default constructor.
Definition at line 29 of file NeutrinoIdTool.cc.
|
private |
Collect all 2D hits in a supplied list of Pfos and push them on to an existing hit list, check so not to double count.
| pfos | input list of pfos |
| reconstructedCaloHitList | output list of all 2d hits in the input pfos |
| reconstructableCaloHitSet | set of reconstructable calo hits |
Definition at line 166 of file NeutrinoIdTool.cc.
References lar_content::LArPfoHelper::GetCaloHits().
Referenced by lar_content::NeutrinoIdTool< T >::GetBestMCSliceIndex().
|
private |
Count the number of neutrino induced hits in a given list using MC information.
| caloHitSet | input list of calo hits |
Definition at line 188 of file NeutrinoIdTool.cc.
References lar_content::LArMCParticleHelper::GetParentMCParticle(), and lar_content::LArMCParticleHelper::IsNeutrino().
Referenced by lar_content::NeutrinoIdTool< T >::GetBestMCSliceIndex().
|
private |
Get the slice with the most neutrino induced hits using Monte-Carlo information.
| pAlgorithm | address of the master algorithm |
| nuSliceHypotheses | the input neutrino slice hypotheses |
| crSliceHypotheses | the input cosmic slice hypotheses |
| bestSliceIndex | the index of the slice with the most neutrino hits |
Definition at line 98 of file NeutrinoIdTool.cc.
References lar_content::NeutrinoIdTool< T >::Collect2DHits(), lar_content::NeutrinoIdTool< T >::CountNeutrinoInducedHits(), f, lar_content::LArMCParticleHelper::GetMCPrimaryMap(), lar_content::LArMCParticleHelper::PrimaryParameters::m_maxPhotonPropagation, lar_content::LArMCParticleHelper::PrimaryParameters::m_selectInputHits, lar_content::NeutrinoIdTool< T >::PassesQualityCuts(), and lar_content::LArMCParticleHelper::SelectCaloHits().
Referenced by lar_content::NeutrinoIdTool< T >::SelectOutputPfos().
|
private |
Use the current MCParticle list to get the nuance code of the neutrino in the event.
| pAlgorithm | address of the master algorithm |
Definition at line 209 of file NeutrinoIdTool.cc.
References lar_content::LArMCParticleHelper::GetNuanceCode(), and lar_content::LArMCParticleHelper::GetTrueNeutrinos().
Referenced by lar_content::NeutrinoIdTool< T >::PassesQualityCuts().
|
private |
Get the features of each slice.
| pTool | the address of the this NeutrinoId tool |
| nuSliceHypotheses | the input neutrino slice hypotheses |
| crSliceHypotheses | the input cosmic slice hypotheses |
| sliceFeaturesVector | vector to hold the slice features |
Definition at line 88 of file NeutrinoIdTool.cc.
Referenced by lar_content::NeutrinoIdTool< T >::SelectOutputPfos().
|
private |
Determine if the event passes the selection cuts for training and has the required NUANCE code.
| pAlgorithm | address of the master algorithm |
| purity | purity of best slice |
| completeness | completeness of best slice |
Definition at line 153 of file NeutrinoIdTool.cc.
References lar_content::NeutrinoIdTool< T >::GetNuanceCode(), lar_content::NeutrinoIdTool< T >::m_minCompleteness, lar_content::NeutrinoIdTool< T >::m_minPurity, lar_content::NeutrinoIdTool< T >::m_nuance, and lar_content::NeutrinoIdTool< T >::m_selectNuanceCode.
Referenced by lar_content::NeutrinoIdTool< T >::GetBestMCSliceIndex().
|
private |
Definition at line 594 of file NeutrinoIdTool.cc.
References lar_content::LArFileHelper::FindFileInPath(), lar_content::NeutrinoIdTool< T >::m_defaultProbability, lar_content::NeutrinoIdTool< T >::m_filePathEnvironmentVariable, lar_content::NeutrinoIdTool< T >::m_maxNeutrinos, lar_content::NeutrinoIdTool< T >::m_minCompleteness, lar_content::NeutrinoIdTool< T >::m_minProbability, lar_content::NeutrinoIdTool< T >::m_minPurity, lar_content::NeutrinoIdTool< T >::m_mva, lar_content::NeutrinoIdTool< T >::m_nuance, lar_content::NeutrinoIdTool< T >::m_persistFeatures, lar_content::NeutrinoIdTool< T >::m_selectNuanceCode, lar_content::NeutrinoIdTool< T >::m_trainingOutputFile, and lar_content::NeutrinoIdTool< T >::m_useTrainingMode.
|
private |
Select all pfos under the same hypothesis.
| pAlgorithm | address of the master algorithm |
| hypotheses | the lists of slices under a certain hypothesis |
| selectedPfos | the list of pfos to populate |
Definition at line 229 of file NeutrinoIdTool.cc.
References lar_content::NeutrinoIdTool< T >::SelectPfos().
Referenced by lar_content::NeutrinoIdTool< T >::SelectOutputPfos().
|
virtual |
Select which reconstruction hypotheses to use; neutrino outcomes or cosmic-ray muon outcomes for each slice.
| pAlgorithm | the address of the master instance, used to access MCParticles when in training mode |
| nuSliceHypotheses | the parent pfos representing the neutrino outcome for each slice |
| crSliceHypotheses | the parent pfos representing the cosmic-ray muon outcome for each slice |
| sliceNuPfos | to receive the list of selected pfos |
Implements lar_content::SliceIdBaseTool.
Definition at line 46 of file NeutrinoIdTool.cc.
References lar_content::NeutrinoIdTool< T >::GetBestMCSliceIndex(), lar_content::NeutrinoIdTool< T >::SliceFeatures::GetFeatureVector(), lar_content::NeutrinoIdTool< T >::GetSliceFeatures(), lar_content::NeutrinoIdTool< T >::m_trainingOutputFile, lar_content::NeutrinoIdTool< T >::m_useTrainingMode, lar_content::LArMvaHelper::ProduceTrainingExample(), lar_content::NeutrinoIdTool< T >::SelectAllPfos(), and lar_content::NeutrinoIdTool< T >::SelectPfosByProbability().
|
private |
Add the given pfos to the selected Pfo list.
| pfos | the pfos to select |
| selectedPfos | the list of pfos to populate |
Definition at line 321 of file NeutrinoIdTool.cc.
Referenced by lar_content::NeutrinoIdTool< T >::SelectAllPfos(), and lar_content::NeutrinoIdTool< T >::SelectPfosByProbability().
|
private |
Select pfos based on the probability that their slice contains a neutrino interaction.
| pAlgorithm | address of the master algorithm |
| nuSliceHypotheses | the input neutrino slice hypotheses |
| crSliceHypotheses | the input cosmic slice hypotheses |
| sliceFeaturesVector | vector holding the slice features |
| selectedPfos | the list of pfos to populate |
Definition at line 247 of file NeutrinoIdTool.cc.
References lar_content::NeutrinoIdTool< T >::m_defaultProbability, lar_content::NeutrinoIdTool< T >::m_maxNeutrinos, lar_content::NeutrinoIdTool< T >::m_minProbability, lar_content::NeutrinoIdTool< T >::m_mva, lar_content::NeutrinoIdTool< T >::m_persistFeatures, lar_content::NeutrinoIdTool< T >::SelectPfos(), and value.
Referenced by lar_content::NeutrinoIdTool< T >::SelectOutputPfos().
|
private |
Default probability set if score could not be calculated.
Definition at line 269 of file NeutrinoIdTool.h.
Referenced by lar_content::NeutrinoIdTool< T >::ReadSettings(), and lar_content::NeutrinoIdTool< T >::SelectPfosByProbability().
|
private |
The environment variable providing a list of paths to mva files.
Definition at line 275 of file NeutrinoIdTool.h.
Referenced by lar_content::NeutrinoIdTool< T >::ReadSettings().
|
private |
The maximum number of neutrinos to select in any one event.
Definition at line 270 of file NeutrinoIdTool.h.
Referenced by lar_content::NeutrinoIdTool< T >::ReadSettings(), and lar_content::NeutrinoIdTool< T >::SelectPfosByProbability().
|
private |
Minimum completeness of the best slice to use event for training.
Definition at line 265 of file NeutrinoIdTool.h.
Referenced by lar_content::NeutrinoIdTool< T >::PassesQualityCuts(), and lar_content::NeutrinoIdTool< T >::ReadSettings().
|
private |
Minimum probability required to classify a slice as the neutrino.
Definition at line 268 of file NeutrinoIdTool.h.
Referenced by lar_content::NeutrinoIdTool< T >::ReadSettings(), and lar_content::NeutrinoIdTool< T >::SelectPfosByProbability().
|
private |
Minimum purity of the best slice to use event for training.
Definition at line 264 of file NeutrinoIdTool.h.
Referenced by lar_content::NeutrinoIdTool< T >::PassesQualityCuts(), and lar_content::NeutrinoIdTool< T >::ReadSettings().
|
private |
The mva.
Definition at line 274 of file NeutrinoIdTool.h.
Referenced by lar_content::NeutrinoIdTool< T >::ReadSettings(), and lar_content::NeutrinoIdTool< T >::SelectPfosByProbability().
|
private |
Nuance code to select for training.
Definition at line 263 of file NeutrinoIdTool.h.
Referenced by lar_content::NeutrinoIdTool< T >::PassesQualityCuts(), and lar_content::NeutrinoIdTool< T >::ReadSettings().
|
private |
If true, the mva features will be persisted in the metadata.
Definition at line 272 of file NeutrinoIdTool.h.
Referenced by lar_content::NeutrinoIdTool< T >::ReadSettings(), and lar_content::NeutrinoIdTool< T >::SelectPfosByProbability().
|
private |
Should select training events by nuance code.
Definition at line 262 of file NeutrinoIdTool.h.
Referenced by lar_content::NeutrinoIdTool< T >::PassesQualityCuts(), and lar_content::NeutrinoIdTool< T >::ReadSettings().
|
private |
Output file name for training examples.
Definition at line 261 of file NeutrinoIdTool.h.
Referenced by lar_content::NeutrinoIdTool< T >::ReadSettings(), and lar_content::NeutrinoIdTool< T >::SelectOutputPfos().
|
private |
Should use training mode. If true, training examples will be written to the output file.
Definition at line 260 of file NeutrinoIdTool.h.
Referenced by lar_content::NeutrinoIdTool< T >::ReadSettings(), and lar_content::NeutrinoIdTool< T >::SelectOutputPfos().